Researchers at the University of Illinois at Urbana-Champaign and at Howard Hughes Medical Institute,
both in Urbana, have the beat — at least on the nanometer scale. They have
constructed a DNA-based nanomechanical device that displays an adjustable ticking
rate, somewhat like a metronome. The rate can be changed by adjusting ion concentration
or by altering a set of DNA-based deactivating and activating switches. The latter
may hold the key to a single-molecule sensor that can detect small sequence differences
in target DNA.
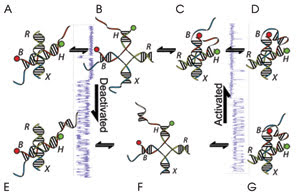
esearchers have developed a stochastic nanoscale
metronome using DNA constructs that flip between two conformations (A and D), modifying
the time spent in each state by attaching single-strand DNA to the construct. They
modified the metronomic cycling rate by adding a deactivator (E) and an activator
(F). Courtesy of Taekjip Ha.
“We showed that even a single basepair difference
can be detected by the device,” said Taekjip Ha, an associate professor of
physics at the university and an investigator at the institute. He led the team
that fabricated the nano-metronome.
The genesis of the device was the group’s
research on four-way Holliday junctions. These DNA constructs are tetrahedral, with
strands that can be exchanged, creating a structure that naturally “ticks”
from one conformation to another. In building the nanometronome, the scientists
combined this function with zipping and unzipping via single-strand DNA.
Measuring conformations
The group started by selecting a junction where
the lifetimes of the two conformers were about equal. It added two single-stranded
overhangs on the end of the sequence: In one conformation, these overhangs were
far from other structures and had little impact; in the other, they fell near other
parts of the structure and, in effect, created a sticky force that kept the junction
in that conformation longer.
To determine the state of the metronome,
the researchers used a total internal reflection fluorescence setup — comprising
an Olympus inverted microscope equipped with a frequency-doubled Nd:YAG 532-nm laser
supplied by CrystaLaser of Reno, Nev. — to study Förster resonance energy
transfer in the system. The distance between the acceptor and donor molecules went
from far to near and back again as the junction flipped between conformations. When
this happened, the fluorescence of the two changed, leading to an observable fluctuation.
The investigators used total internal
reflection fluorescence microscopy because it reduces background fluorescence and
provides excellent signal to noise in single-molecule fluorescence measurements,
according to Ha. Because the imaging had to be done at high speed and low light
levels, the researchers also used an electron-multiplying CCD camera from Andor
Technology of Belfast, UK.
Ha noted that a traditional metronome
ticks regularly and periodically, but that the nanoscale version did not change
state at a given frequency. Instead, it flipped between states at a variable rate.
The investigators controlled that rate
by two means. The first was by altering the concentration of doubly charged magnesium
ions, which were critical for the occurrence of conformational transitions. Changing
the concentration directly also affected the speed of the changes.
The second was by introducing a short,
single-stranded deactivator/activator. The deactivator would bind to one of the
free overhangs, rendering it inactive and thereby decreasing the tendency for the
junction to prefer a particular conformation. Subsequent addition of an activator
strand removed the deactivator and restored the metronome’s rate.
In tests, the researchers could see
a difference in the rate if the overhangs differed by as little as one base pair,
indicating that the technique potentially could be used as a single-molecule DNA
sensor, which Ha noted is the subject of ongoing research.
“We are designing constructs
that can detect single-strand DNA in solution, and it should work at the research/publication
level,” he said.
Nano Letters, March 8, 2006, pp. 496-500.